Cytkit | Honeychrome
 Honeychrome
Honeychrome
- Overview and Installation
- Acquire Data
- Spectral Analysis
- AutoSpectral in Honeychrome
- Conventional Analysis
- Manipulate Plots and Gates
- Reports, Exports & Sample Comparison
- Programming and Plugins
AutoSpectral in Honeychrome
AutoSpectral is a set of tools to enhance spectral analysis by providing you with the best possible spectral signatures of your spectral controls, optimising the removal of autofluorescence, and providing reproducible mixing. See the Colibri Cytometry blog and the AutoSpectral Github repository for more information on AutoSpectral.
Honeychrome now provides a tab for AutoSpectral AF (the per-cell autofluorescence extraction part of AutoSpectral), which radically clears up artifacts due to variable autofluorescence, for example negative spreading. See AutoSpectral AF on the Colibri Cytometry blog
How to perform AutoSpectral AF (per-cell autofluorescence extraction)
This guide assumes you have already installed Honeychrome using the executable installers (Windows/Mac/Linux) or Python (cross-platform), that you have a set of FCS files to import, and that you are already familiar with the basic spectral unmixing process. If not, please go to Spectral Analysis first.
We will be continuing use of AutoSpectral Full Workflow Example (a 9-colour panel from the 5-laser Cytek Aurora from the AutoSpectral Full Workflow Example as set up in the tutorial Spectral Analysis.
Show the AutoSpectral AF process
Go to the AutoSpectral tab and check the box “Show AutoSpectral AF process”
Extract AF profile from an unstained sample
Under Step 1, select your (first) unstained sample, and press the button Extract AF Profile. (Default number of clusters: 100.) You should see a set of spectra of clusters below. This step should be repeated for each unstained control that you have available, generating multiple AF profiles. After extraction, the clusters can be inspected by clicking on the Stored AF Profiles list.
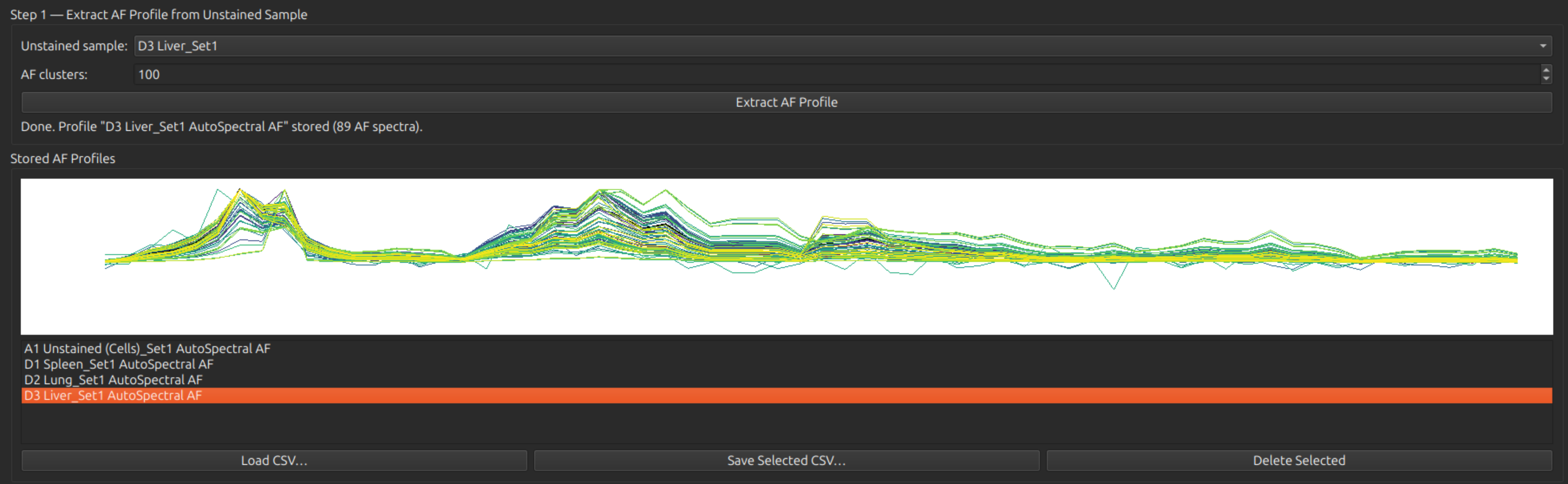
Assign the AF profiles to samples
Under Step 2, select the AF profiles to apply to each sample. Multiple controls (columns) can be selected for each sample (row). Each cell in the sample is automatically assigned its nearest cluster from of the selected controls for a more accurate AF subtraction.
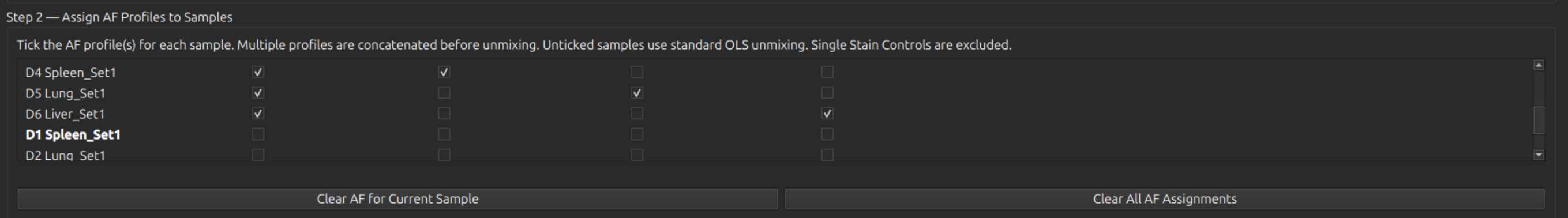
Inspect the AutoSpectral AF results against standard unmixing
Under Step 3, select a sample (from the sample browser, left pane), and select channels and a source gate on the side-by-side plots.
Negative spread and other artifacts caused by inhomogeneous autofluorescence should be much improved.
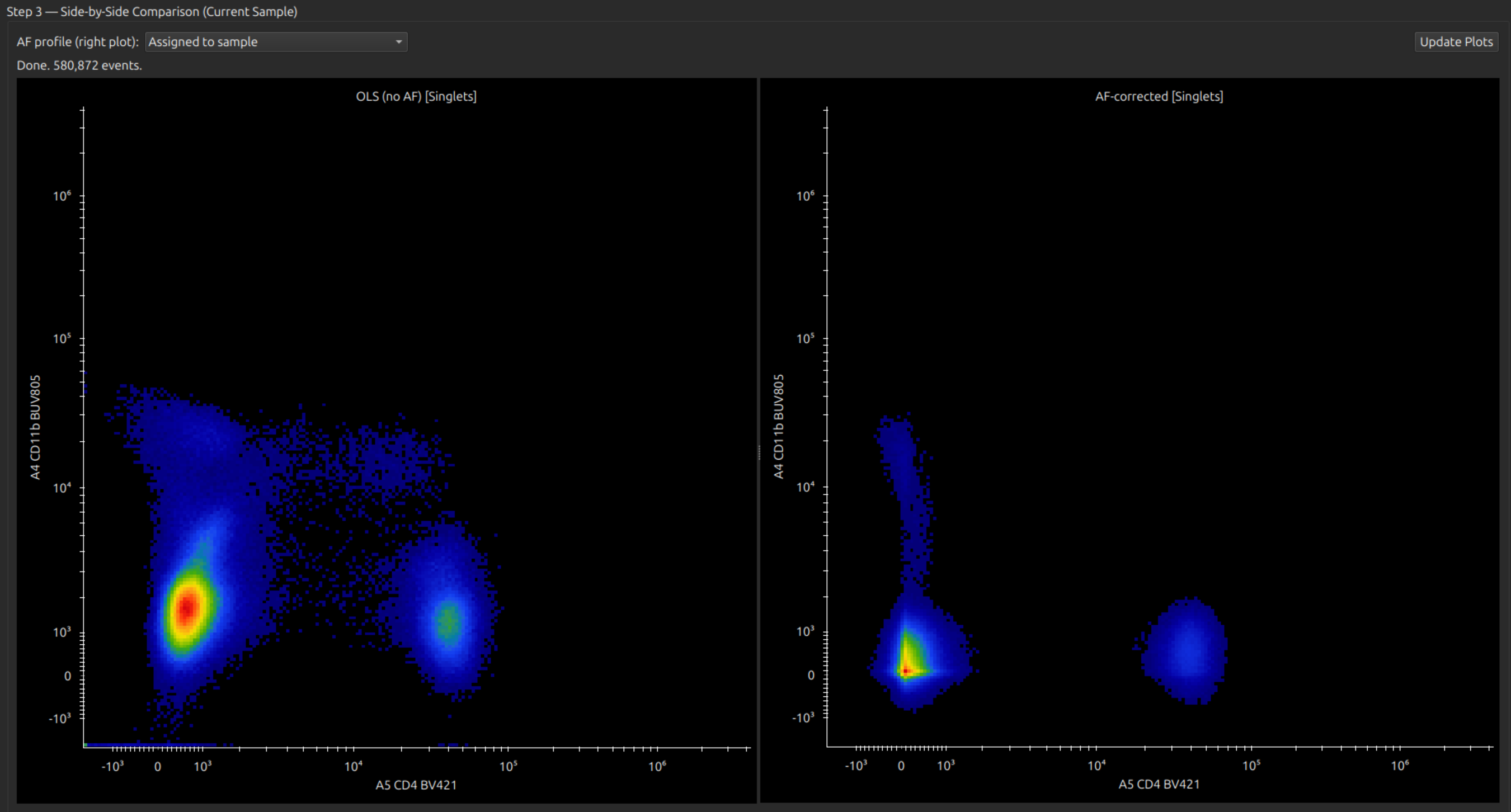
This workflow recalculates all unmixed data for analysis in the Unmixed Data / Statistics tabs, and is applied whenever an assigned sample is loaded in the sample browser.