Cytkit | Honeychrome
 Honeychrome
Honeychrome
- Overview and Installation
- Acquire Data
- Spectral Analysis
- AutoSpectral in Honeychrome
- Conventional Analysis
- Manipulate Plots and Gates
- Reports, Exports & Sample Comparison
- Programming and Plugins
Programming and Plugins
Honeychrome can be extended with Python plugins that define a new tab in the Honeychrome main window. The guide below specifies how to set one up and gives examples.
Plugins have full access to the data in the application:
- the experiment’s metadata (raw and unmixed cytometry, spectral process, list of samples and controls, settings)
- the ephemeral data (gating hierarchy, transforms, lookup tables, current plots, etc)
- the current sample
- the set of signals to communicate with other parts of the GUI
Note: If you want to use the plugin functionality, please run Honeychrome in Python: see the installation page for instructions.
Specification of a Plugin
A plugin must be named *_tab.py and placed in the Experiments/plugins folder (within the user’s home folder) to be found by Honeychrome. The minimal plugin must provide a name (‘plugin_name’), which will be used as the tab’s name, and a class PluginWidget (subclass of PySide6.QtWidgets.QWidget), which will be displayed within the tab.
The initialisation method of PluginWidget must accept the following arguments:
- controller: this object contains all data in the application
- bus: the signals bus which can be used to communicate with the rest of the GUI
The minimal code for a plugin is the following:
from PySide6.QtWidgets import QWidget
plugin_name = 'Blank Example Plugin'
class PluginWidget(QWidget):
"""
The main UI container for the plugin.
Required arguments:
bus: the signals to communicate with the rest of the honeychrome app
controller: the honeychrome controller including all ephemeral data and the experiment model
"""
def __init__(self, bus=None, controller=None, parent=None):
super().__init__(parent)
self.bus = bus
self.controller = controller
Use the FlowKit objects through Honeychrome
FlowKit is a Python toolkit for flow cytometry analysis and visualization, which has excellent documentation and tutorials for programmers. FlowKit handles sample data, FCS file input/output, transforms, gates and gating strategies, sessions and workspaces.
Honeychrome uses the FlowKit objects for the sample data, channel transforms and specification of gating strategies in the raw and unmixed data. These can be accessed through the controller:
controller.current_sample # current sample (flowkit.Sample object)
controller.raw_transformations # transformations of the data in all raw channels (wrapper to flowkit.transform objects)
controller.unmixed_transformations # transformations of the data in all raw channels (wrapper to flowkit.transform objects)
controller.raw_gating # gating hierarchy for raw data (flowkit.GatingStrategy object)
controller.unmixed_gating # gating hierarchy for unmixed data (flowkit.GatingStrategy object)
Note that controller.raw_transformations and controller.unmixed_transformations are dictionaries of the current flowkit.transforms objects assigned to each channel, which are accessed with controller.raw_transformations[raw_channel_name].xform or controller.unmixed_transformations[unmixed_label_name].xform, yielding a flowkit.LogicleTransform, flowkit.LinearTransform or flowkit.LogTransform object, or None if not assigned.
Accessing Data
The following data can be accessed through the controller:
- Experiment object (data saved in the .kit file) contains all data necessary to re-generate the state of the controller (along with the sample data in the FCS files)
controller.experiment.settings # settings controller.experiment.samples # list of samples controller.experiment.process # data associated with the spectral process, including the spectral model, profiles, spillover matrix and AutoSpectral AF profiles controller.experiment.cytometry # data associated with raw and unmixed cytometry controller.experiment.statistics # statistical comparison data
See experiment_model.py for a full definition.
- Ephemeral data (data in memory of the running application)
controller.experiment_dir # absolute path of current experiment folder
controller.current_sample # current sample (flowkit.Sample object)
controller.current_sample_path # path of current sample (relative to experiment folder)
controller.live_sample_path # path of live acquisition sample data (relative to experiment folder)
controller.raw_event_data # numpy array of raw sample data of current sample in all channels
controller.unmixed_event_data # numpy array of unmixed sample data of current sample in all channels
controller.transfer_matrix # matrix that transforms the raw data into the unmixed data (i.e. the unmixing matrix, plus extra rows/columns for the non-fluorescence channels
controller.raw_transformations # transformations of the data in all raw channels (wrapper to flowkit.transform objects)
controller.unmixed_transformations # transformations of the data in all raw channels (wrapper to flowkit.transform objects)
controller.raw_gating # gating hierarchy for raw data (flowkit.GatingStrategy object)
controller.unmixed_gating # gating hierarchy for unmixed data (flowkit.GatingStrategy object)
controller.raw_lookup_tables # lookup tables for fast gating in all gates defined on raw data
controller.unmixed_lookup_tables # lookup tables for fast gating in all gates defined on unmnixed data
controller.current_mode # name of tab currently live in the main window
controller.data_for_cytometry_plots_raw # a copy of the cytometry_data_dictionary for raw data
controller.data_for_cytometry_plots_unmixed # a copy of the cytometry_data_dictionary for unmixed data
The cytometry data dictionary bundles ephemeral data that is required for cytometry plots or statistical comparisons, and is defined as follows:
cytometry_data_dictionary = {
'pnn': None, # list of channel names
'fluoro_indices': None, # list of fluorescence channel indices to the list of channel names
'lookup_tables': None, # dictionary of boolean lookup tables for each gate on the 1D or 2D plot on which the gate is defined (for fast gating)
'event_data': None, # event data (which may be raw or unmixed depending on the copy of the dictionary)
'transformations': None, # set of transforms for all channels
'statistics': {}, # event statistics for each gate in the hierarchy
'gating': GatingStrategy(), # flowkit.GatingStrategy object used to define the gating lookup tables
'plots': [], # set of cytometry plot definitions (1D histograms, 2D histograms, ribbon plots referencing the channel names, source gates and child gates
'histograms': [], # set of 1D and 2D histograms for plotting on the plots
'gate_membership': {} # dictionary of gate membership for each gate, boolean array corresponding to event_data
}
The controller also contains the communications to the instrument driver and trace analyser processes, and the live queue of oscilloscope traces.
The full definition of the controller is according to the source code.
The transform class is a wrapper around the flotkit.Transform, as defined in the source code.
Reusable functions and classes
Honeychrome contains many reusable functions and classes that may be useful in a plugin. See the following in particular, which are all demonstrated in the data_processing_example_tab.py
ExportablePlotWidget
ExportablePlotWidget puts a matplotlib figure into a widget, with a button for export. Arguments are:
- figure (required): a matplotlib figure object
- title (optional): a string for the figure title, also used as the filename on export
- experiment_dir (optional): absolute path of folder to save the image
Example Python code:
from honeychrome.view_components.exportable_plot_widget import ExportablePlotWidget
from matplotlib import pyplot as plt
figure, ax = plt.subplots(1)
ax.scatter(np.arange(100), np.arange(100)**2)
plot_widget = ExportablePlotWidget(figure, title="Example Plot", experiment_dir=self.controller.experiment_dir)
Example display of this widget:
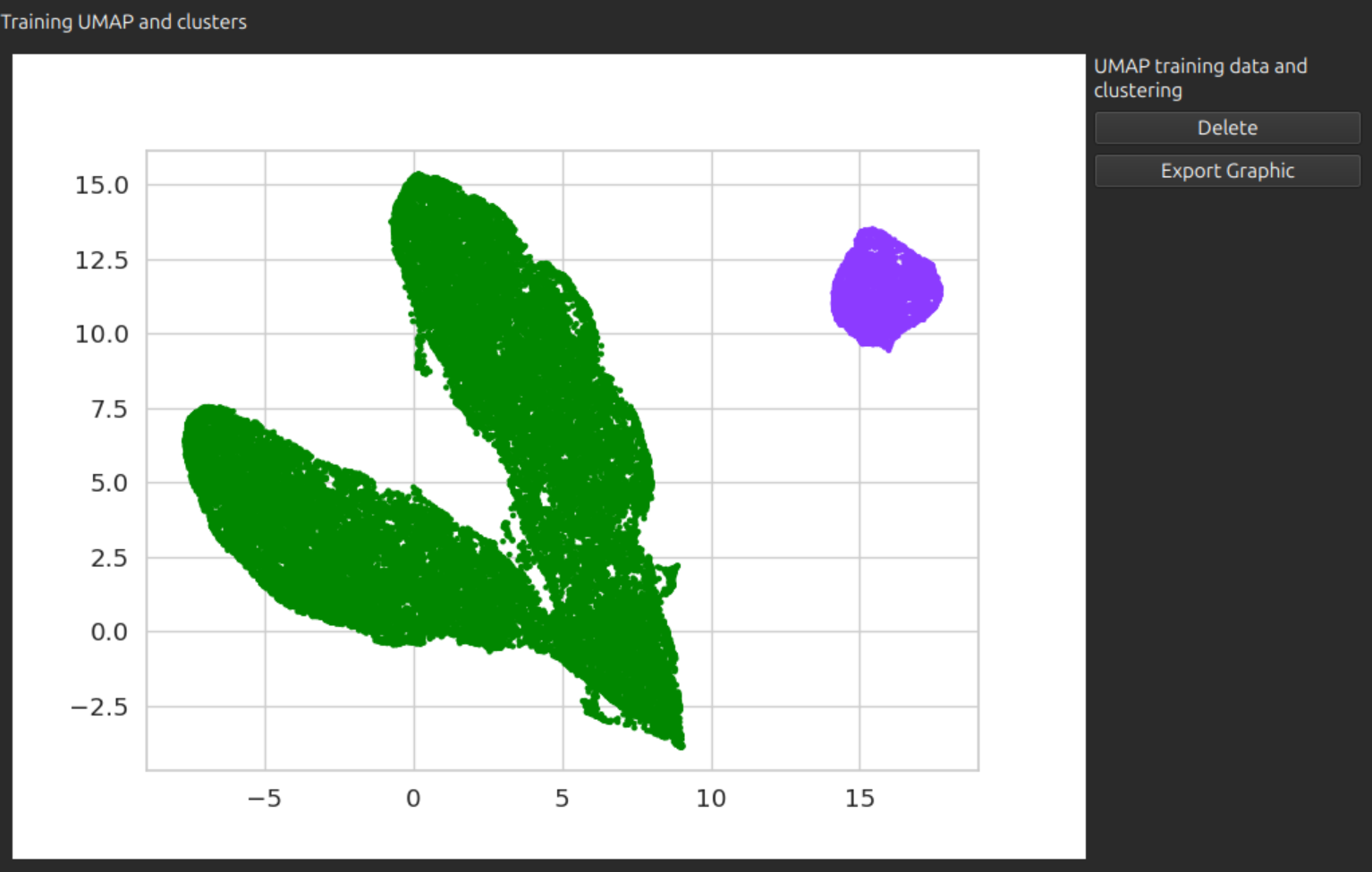
OrderedMultiSamplePicker
OrderedMultiSamplePicker is a sample picker widget, which produces an ordered list of samples as an output. Arguments are:
- title (optional): string
- source_samples (optional): list of source samples
The list of source samples can be updated after initialisation with the method OrderedMultiSamplePicker.set_items(source_samples)
Example Python code:
- set up widget
from honeychrome.view_components.ordered_multi_sample_picker import OrderedMultiSamplePicker self.picker = OrderedMultiSamplePicker(title="Choose Source Samples for Processing") - add samples to it
from pathlib import Path all_samples = self.controller.experiment.samples['all_samples'] source_samples_relative_to_raw = [str(Path(sample).relative_to(self.controller.experiment.settings['raw']['raw_samples_subdirectory'])) for sample in all_samples] self.picker.set_items(source_samples_relative_to_raw) - retrieve selection
selection = self.picker.get_ordered_list()
Example display of this widget:
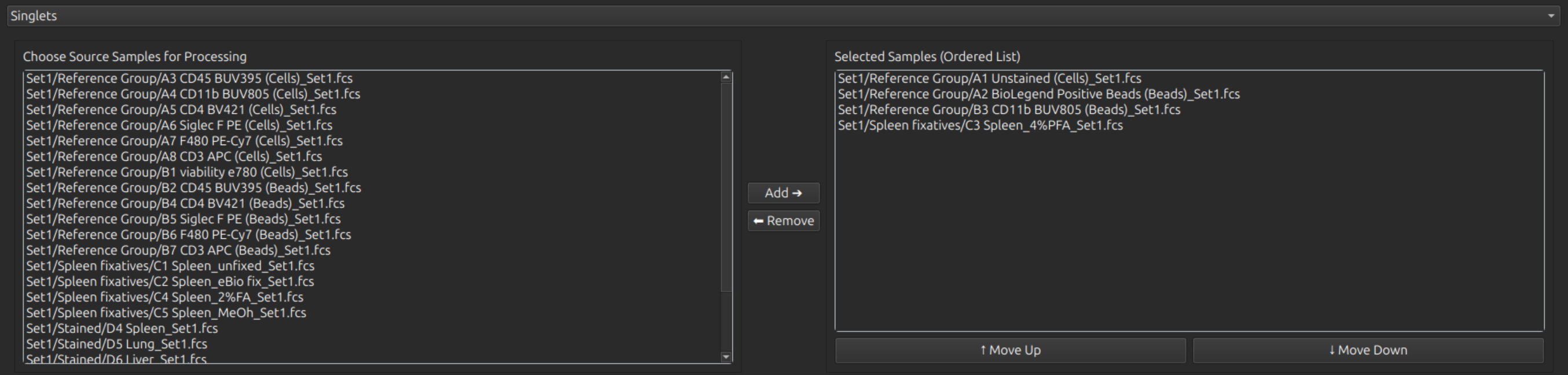
CopyableTableWidget
CopyableTableWidget displays a table which can be copy/pasted e.g. into a spreadsheet. Arguments:
- list_of_dicts: list where each element is a dict containing the data for a row of the table
- headers: ordered list of columns (strings)
Example Python code:
from honeychrome.view_components.copyable_table_widget import CopyableTableWidget
headers = ['Index', 'Colour', 'Count']
list_of_dicts = [
{'Index': -1, 'Colour': '#7f7f7f', 'Count': 0},
{'Index': 0, 'Colour': '#8c3bff', 'Count': 3691},
{'Index': 1, 'Colour': '#018700', 'Count': 36414}
]
table_widget = CopyableTableWidget(table_data, table_headers)
Example display of this widget:
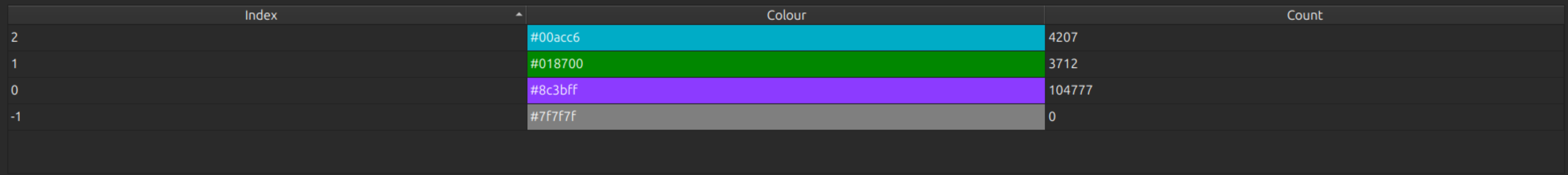
with_busy_cursor
Use as a decorator on any heavy function or class method so that a mouse spinner is displayed while running.
Example Python code:
from honeychrome.view_components.busy_cursor import with_busy_cursor
@with_busy_cursor
def example_function():
...
Example Plugins
Two plugins are provided in the Honeychrome package as examples:
- Hello World Example Plugin hello_world_example_plugin_tab.py
- demonstrates the minimal plugin
- Data Processing Example Plugin data_processing_example_plugin_tab.py
- demonstrates a plugin that accesses unmixed data over a set of samples, displays sample picker, demonstrates a toy UMAP workflow, produces graphs, tables, output text
These are automatically copied to the Experiments/plugins folder when Honeychrome is started. To enable the plugins, go to menu Edit > App Configuration.